 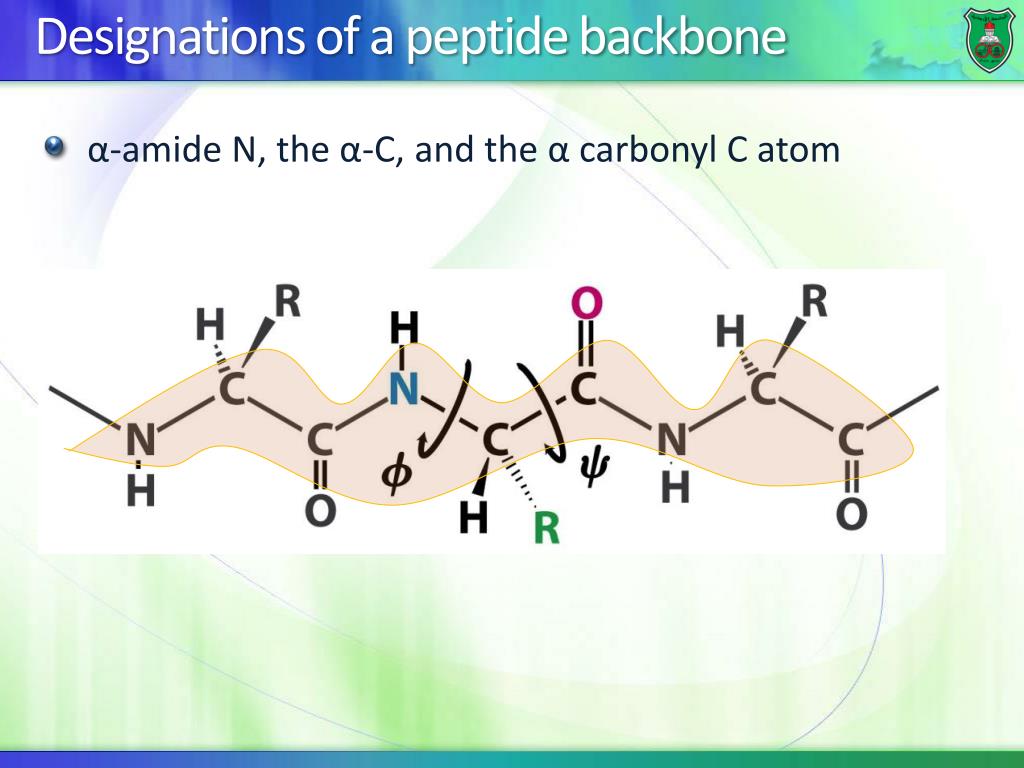
Up to which layer is there a degree of localization? This distance is related with the localization of the acyl-chains around the proteinĪt which distance from the protein is the first layer of lipids located? You will see the distance distribution of the C40 atom ortogonal to the protein. Right click pressed on that set and then -> "Hide"Īt the right (Destination), select Graph G1(0 sets) In the new window, right click pressed on G0.S0-> Selector operations -> Select AllĬhange Operation type from "Sort" to "Join"Īt the left (Source), click on Set: G0.S0 Create an index group with the C40 atoms (located in the middle of the one of the acyl chains) and split that group into residues: Let us check how the lipids accommodate around the protein. Therefore it is an ideal system to study lipid-protein interactions. It has been possible to crystallize it with a complete layer of lipids around it. Localization of lipids around membrane proteinsĪquaporin-0 has a unique feature. For a detail analysis of the permeation of water through aquaporins we recomend to read the literature below and to follow the tutorial developed by Prof. There are other aquaporins which exhibit a more high water permeability, while still maintaing strict control on the substances that cross. In the created representation change the Selected Atoms to "resid 525 9370"įor that representation, choose VDW as Drawing MethodĪquaporin-0 is a poorly conducting channel. In the vmd Graphical Representations window:Ĭlick on the First representation of the list to highlight it.Ĭlick on "Create Rep". Highlight two water molecules to evidence this. However, water molecules by sometimes manage to diffuse trhough the pore. 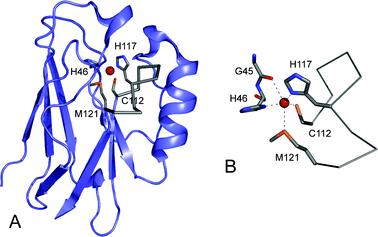
Under equilibrium conditions, the net flux of water is zero. From the trajectory, could you imagine how these residues facilitate the water conduction through such narrow pores? HINT: look whether hydrogen bonds may be formed between these residues and water. We selected key residues at the pore of each aquaporin tetramer. What is the advantage to allow the water permeation in a single-file fashion? How would you describe the water passage through each one of these paths? How many water pathways can you identify? Also fraphical representations can be hidden/shown for more clarity. In the VMD Main window, press play, adjusting the speed, orientation and zoom. Replace "resname SOL and within 4 of protein" by "protein and resid 62 64 180 183" Look at the protein with pymolįor "Selected Atoms" choose "resname SOL and within 4 of protein" Select "PDB Format" and save the file in the Desktop directory. Download the structure, by clicking on the "Download Files" button at the right side. Please go to the protein data bank, search the structure with PDB code 2B6O. We will consider aquaporin-0, the most abundant protein in our eye lense. We will consider an aquaporin, a class of membrane proteins which facilitate the permeation of water and other solutes through biological lipid bilayers in response to osmotic pressure. Now, we will setup an MD simulation of a membrane protein. What change in conformation (area per lipid and thickness), ordering, and dynamics (diffusion coefficient) is expected if the temperature is reduced?Ĭ. What is the origin of the discrepancies between the simulation and the experimental value? How does the estimated value compares with the experimental diffusion coefficient for this lipid at 320 K (D exp~1.5x10 -7 cm 2/s)? The slope of this curve is related to the diffusion coefficient. Which type of diffusive behavior do the lipids display at that given time scale (sub-diffusive, diffusive or super-diffusive)?. Use the zoom button at the up-left xmgrace menu for this purpose. Zoom to the time window, considered for the fitting (0 to 10000 ps). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
